Distributions
This section introduces the computation of Recurrence Microstates Analysis (RMA) distributions using RecurrenceMicrostatesAnalysis.jl.
We begin with a Quick start with RecurrenceMicrostatesAnalysis.jl, which demonstrates a simple application example. Next, we present A brief review of Recurrence Plots (RP) and RMA. Finally, we explain the distribution function in Computing RMA distributions, including the role of Histograms.
Quick start with RecurrenceMicrostatesAnalysis.jl
This section presents concise examples illustrating how to use the package. RMA distributions are computed using the distribution function, which returns a Probabilities structure containing the microstate distribution.
We start with a simple example based on a uniform random process. First, we generate the data and convert it into a StateSpaceSet:
using Distributions, RecurrenceMicrostatesAnalysis
data = rand(Uniform(0, 1), 10_000);
ssset = StateSpaceSet(data)1-dimensional StateSpaceSet{Float64} with 10000 points
0.28769478368805634
0.3270312001740592
0.8762853878338924
0.022373842486420492
0.6444688570573011
0.3995735932242024
0.43763516491971943
0.24545837329781994
0.1739045528719606
0.8475601713614623
⋮
0.15006206550700574
0.35127293917374447
0.823147175449717
0.9025771188676037
0.6613568435448977
0.41021813200489277
0.981946572913893
0.18291558665176633
0.6353629851762662Next, we compute the RMA distribution. This requires specifying the recurrence threshold $\varepsilon$ and the microstate size $N$. These parameters are discussed in more detail in A brief review and Optimizing a parameter.
ε = 0.27
N = 2
dist = distribution(ssset, ε, N) Probabilities{Float64,1} over 16 outcomes
1 0.11082574298344809
2 0.047184227408636245
3 0.04726184291621466
4 0.08552848859202068
5 0.04713421741663985
6 0.08556669622590593
7 0.07679594382957715
8 0.030660525973089423
9 0.047239838519736246
10 0.0769313708879034
11 0.08582194722505557
12 0.030578109506279354
13 0.08565031293252391
14 0.030441482208145187
15 0.030611316140964965
16 0.08176793723385932As another example, we use DynamicalSystems.jl to generate data from the Hénon map, following the example presented in its documentation:
using DynamicalSystems
function henon_rule(u, p, n) # here `n` is "time", but we don't use it.
x, y = u # system state
a, b = p # system parameters
xn = 1.0 - a*x^2 + y
yn = b*x
return SVector(xn, yn)
end
u0 = [0.2, 0.3]
p0 = [1.4, 0.3]
henon = DeterministicIteratedMap(henon_rule, u0, p0)
total_time = 10_000
X, t = trajectory(henon, total_time)
X2-dimensional StateSpaceSet{Float64} with 10001 points
0.2 0.3
1.244 0.06
-1.10655 0.3732
-0.341035 -0.331965
0.505208 -0.102311
0.540361 0.151562
0.742777 0.162108
0.389703 0.222833
1.01022 0.116911
-0.311842 0.303065
⋮
-0.582534 0.328346
0.853262 -0.17476
-0.194038 0.255978
1.20327 -0.0582113
-1.08521 0.36098
-0.287758 -0.325562
0.558512 -0.0863275
0.476963 0.167554
0.849062 0.143089Finally, we compute the RMA distribution of the trajectory X. Here, the threshold is selected using optimize by maximizing the recurrence microstate entropy:
ε, S = optimize(Threshold(), RecurrenceEntropy(), X, N)(0.8568661000572908, 2.608204784592693)dist = distribution(X, ε, N) Probabilities{Float64,1} over 16 outcomes
1 0.0512218
2 0.0278322
3 0.0548976
4 0.0533404
5 0.0547514
6 0.0531694
7 0.106138
8 0.0415038
9 0.0155716
10 0.132076
11 0.0652664
12 0.0420656
13 0.0652942
14 0.0418562
15 0.0288536
16 0.1661618A brief review
Recurrence Plots (RPs) were introduced in 1987 by Eckmann et al. (Eckmann et al., 1987) as a method for analyzing dynamical systems through recurrence properties.
Consider a time series $\vec{x}_i \in \mathbb{R}^d$, $i \in \{1, 2, \dots, K\}$, where $K$ is the length of the time series and $d$ is the dimension of the phase space. The recurrence plot is defined by the recurrence matrix
\[R_{i,j} = \Theta(\varepsilon - \|\vec x_i - \vec x_j\|),\]
where $\Theta(\cdot)$ denotes the Heaviside step function and $\varepsilon$ is the recurrence threshold.
The following figure shows examples of recurrence plots for different systems: (a) white noise; (b) a superposition of harmonic oscillators; (c) a logistic map with a linear trend; (d) Brownian motion.
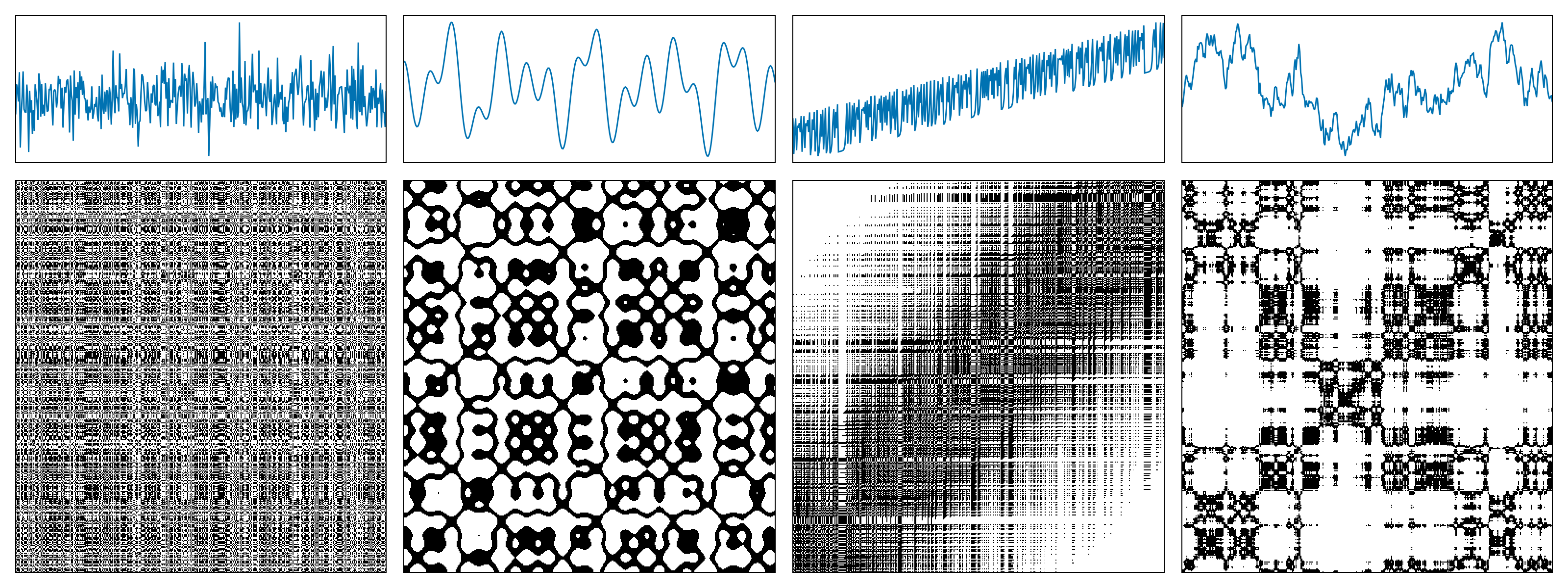
A recurrence microstate is a local structure extracted from an RP. For a given microstate shape and size, the set of possible microstates is finite. For example, square microstates with size $N = 2$ yield $16$ distinct configurations.
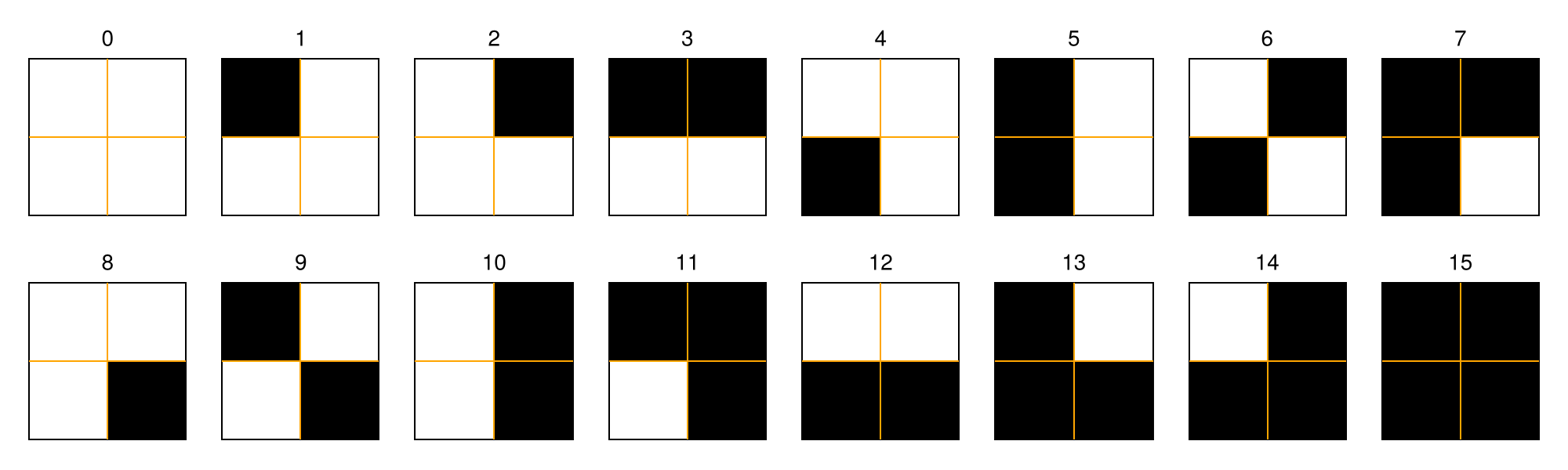
Recurrence Microstates Analysis (RMA) uses the probability distribution of these microstates as a source of information for characterizing dynamical systems.
Computing RMA distributions
The computation of RMA distributions is the core functionality of RecurrenceMicrostatesAnalysis.jl. All other tools in the package rely on these distributions as their primary source of information.
RMA distributions are computed using the distribution function:
RecurrenceMicrostatesAnalysis.distribution — Function
distribution(core::RMACore, [x], [y])Compute an RMA distribution from the recurrence structure constructed using the input data [x] and [y].
If [x] and [y] are identical, the result corresponds to a Recurrence Plot (RP); otherwise, it corresponds to a Cross-Recurrence Plot (CRP).
The core argument must be a subtype of RMACore and defines how the computation is performed, including the execution backend (CPU or GPU), the microstate shape, the recurrence expression, and the sampling mode.
The result is returned as a Probabilities object, where each index corresponds to the decimal representation of the associated microstate.
Internally, distribution calls histogram and converts the resulting counts into probabilities.
distribution(core::CPUCore, [x], [y]; kwargs...)Compute an RMA distribution for the input data [x] and [y] using a CPU backend configuration defined by core, which must be a CPUCore.
For time-series analysis, the inputs [x] and [y] must be provided as StateSpaceSet objects. For spatial analysis, the inputs must be provided as AbstractArrays.
Arguments
core: ACPUCoredefining theMicrostateShape,RecurrenceExpression, andSamplingModeused in the computation.[x]: Input data provided as aStateSpaceSetor anAbstractArray.[y]: Input data provided as aStateSpaceSetor anAbstractArray.
Keyword Arguments
threads: Number of threads used to compute the distribution. By default, this is set toThreads.nthreads(), which can be specified at Julia startup using--threads Nor via theJULIA_NUM_THREADSenvironment variable.
Examples
- Time series:
ssset = StateSpaceSet(rand(Float64, (1000)))
core = CPUCore(Rect(Standard(0.27), 2), SRandom(0.05))
dist = distribution(core, ssset, ssset)- Spatial data:
spatialdata = rand(Float64, (3, 50, 50))
core = CPUCore(Rect(Standard(0.5), (2, 2, 1, 1)), SRandom(0.05))
dist = distribution(core, spatialdata, spatialdata)distribution(core::GPUCore, [x], [y]; kwargs...)Compute an RMA distribution for the input data [x] and [y] using a GPU backend configuration defined by core, which must be a GPUCore.
The inputs [x] and [y] must be vectors of type AbstractGPUVector. This method supports time-series analysis only.
Arguments
core: AGPUCoredefining theMicrostateShape,RecurrenceExpression, andSamplingModeused in the computation.[x]: Input data provided as anAbstractGPUVector.[y]: Input data provided as anAbstractGPUVector.
Keyword Arguments
groupsize: Number of threads per GPU workgroup.
Examples
using CUDA
gpudata = StateSpaceSet(Float32.(data)) |> CuVector
core = GPUCore(CUDABackend(), Rect(Standard(0.27f0; metric = GPUEuclidean()), 2), SRandom(0.05))
dist = distribution(core, gpudata, gpudata)Spatial data are not supported by GPUCore.
A commonly used convenience interface is:
distribution([x], ε::Float, n::Int; kwargs...)This method automatically selects a CPUCore when x is a StateSpaceSet and a GPUCore when x is an AbstractGPUVector. By default, square microstates of size n are used.
Additional keyword arguments include:
rate::Float64: Sampling rate (default:0.05).sampling::SamplingMode: Sampling mode (default:SRandom).metric::Metric: Distance metric from Distances.jl. When using aGPUCore, aGPUMetricmust be provided.
Alternatively, a RecurrenceExpression can be specified directly:
distribution([x], expr::RecurrenceExpression, n::Int; kwargs...)Example:
expr = Corridor(0.05, 0.27)
dist = distribution(ssset, expr, 2) Probabilities{Float64,1} over 16 outcomes
1 0.17799396319384614
2 0.08094677316527843
3 0.08095877556335757
4 0.06394477616627803
5 0.08103439067125612
6 0.06382355194567875
7 0.056042997390878695
8 0.024357466621831043
9 0.08099738327717879
10 0.05581755234695892
11 0.06389956713351327
12 0.0242482447993109
13 0.06402619243324817
14 0.024460087125407655
15 0.024206236406033924
16 0.033242041759943636If a custom MicrostateShape is required, the call simplifies to:
distribution([x], shape::MicrostateShape; kwargs...)Example:
shape = Triangle(Standard(0.27), 3)
dist = distribution(ssset, shape) Probabilities{Float64,1} over 64 outcomes
1 0.03494116947955793
2 0.027660258571376835
3 0.013955379360668395
4 0.013126047793908004
5 0.015479788819570064
6 0.011611242174621414
7 0.02598578911848957
8 0.0235634206555781
9 0.013023406758021857
10 0.01157522777606487
⋮
56 0.006926169082399143
57 0.02146878321953117
58 0.02094277292061366
59 0.007587233375903686
60 0.008456380861068254
61 0.008105240475141962
62 0.007927169282279056
63 0.02031052014595435
64 0.021231288269049967Cross-recurrence plots
RMA distributions can also be computed from Cross-Recurrence Plots (CRPs) by providing two time series:
distribution([x], [y], expr::RecurrenceExpression, n::Int; kwargs...)
distribution([x], [y], shape::MicrostateShape; kwargs...)Example:
data_1 = StateSpaceSet(rand(Uniform(0, 1), 1000))
data_2 = StateSpaceSet(rand(Uniform(0, 1), 2000))
dist = distribution(data_1, data_2, 0.27, 2) Probabilities{Float64,1} over 16 outcomes
1 0.11120569648776678
2 0.04891288019148531
3 0.04990435749266407
4 0.0823326756867733
5 0.048972969724890084
6 0.08534716727924607
7 0.07302881293126759
8 0.03192757208240278
9 0.0499844768705371
10 0.07262821604190244
11 0.08591801784659142
12 0.03148691550410111
13 0.0827533024206067
14 0.031206497681545504
15 0.030926079858989895
16 0.08346436189922986The inputs x and y must have the same phase-space dimensionality. The following example is invalid and will raise an exception:
data_1 = StateSpaceSet(rand(Uniform(0, 1), (1000, 2)))
data_2 = StateSpaceSet(rand(Uniform(0, 1), (2000, 3)))
dist = distribution(data_1, data_2, 0.27, 2)Spatial data
The package also provides experimental support for spatial data, following "Generalised Recurrence Plot Analysis for Spatial Data" (Marwan et al., 2007). In this context, input data are provided as AbstractArrays:
\[ \vec{x}_{\vec i} \in \mathbb{R}^m,\quad \vec{i} \in \mathbb{Z}^d\]
For example:
spatialdata = rand(Uniform(0, 1), (2, 50, 50))2×50×50 Array{Float64, 3}:
[:, :, 1] =
0.0108399 0.132402 0.0573931 0.813676 … 0.905534 0.144873 0.233635
0.989935 0.712167 0.0762763 0.0811511 0.313637 0.730853 0.463623
[:, :, 2] =
0.421137 0.00382678 0.229889 0.405717 … 0.970142 0.501561 0.290975
0.339744 0.971731 0.203877 0.107703 0.846328 0.207389 0.997047
[:, :, 3] =
0.28056 0.501925 0.995227 0.108032 … 0.637218 0.470099 0.151813
0.1607 0.420132 0.924564 0.0215797 0.247394 0.555203 0.421879
;;; …
[:, :, 48] =
0.705112 0.8773 0.753606 0.211476 … 0.506887 0.759426 0.964074
0.586371 0.98676 0.0166541 0.490767 0.166902 0.266415 0.0865137
[:, :, 49] =
0.154691 0.860645 0.901434 0.606442 … 0.930763 0.780562 0.84894
0.528297 0.373878 0.666882 0.00620051 0.997089 0.710053 0.0468884
[:, :, 50] =
0.629583 0.106897 0.415768 0.156681 … 0.353183 0.883065 0.804375
0.309527 0.276529 0.0549814 0.702925 0.555765 0.665665 0.97128Due to the high dimensionality of spatial recurrence plots, direct visualization is often infeasible. RMA distributions provide a compact alternative by sampling microstates directly from the data.
Examples:
- Full $2 \times d$ microstates:
distribution(spatialdata, Rect(Standard(0.27), (2, 2, 2, 2))) Probabilities{Float64,1} over 65536 outcomes
1 0.0524803896739187
2 0.008791254540471341
3 0.008923088665387645
4 0.0023695449294167032
5 0.008871048879236473
6 0.0022030176137329527
7 0.0022966892288050623
8 0.0008048820258047953
9 0.008961251175231838
10 0.002397299482030662
⋮
65528 0.0
65529 0.0
65530 0.0
65531 0.0
65532 0.0
65533 0.0
65534 0.0
65535 0.0
65536 0.0spatialdata_1 = rand(Uniform(0, 1), (2, 50, 50))
spatialdata_2 = rand(Uniform(0, 1), (2, 25, 25))
distribution(spatialdata_1, spatialdata_2, Rect(Standard(0.27), (2, 2, 2, 2))) Probabilities{Float64,1} over 65536 outcomes
1 0.05005133841414915
2 0.00827199236431474
3 0.007867069661166466
4 0.002255997917540384
5 0.007606762209142576
6 0.0024584592691145207
7 0.0027766128215881646
8 0.0008966145569711782
9 0.008055069487628165
10 0.0023283055431025756
⋮
65528 0.0
65529 0.0
65530 0.0
65531 0.0
65532 0.0
65533 0.0
65534 0.0
65535 0.0
65536 0.0- Projected microstates:
distribution(spatialdata, Rect(Standard(0.27), (2, 1, 2, 1))) Probabilities{Float64,1} over 16 outcomes
1 0.4642065805914202
2 0.0951070387338609
3 0.09467388588088296
4 0.02374344023323615
5 0.09461391087047064
6 0.024139941690962098
7 0.021947521865889212
8 0.002838817159516868
9 0.09579341940857976
10 0.022337359433569345
11 0.024203248646397335
12 0.002982090795501874
13 0.023880049979175342
14 0.0028554768846314037
15 0.0029687630154102457
16 0.0037084548104956267distribution(spatialdata_1, spatialdata_2, Rect(Standard(0.27), (2, 1, 2, 1))) Probabilities{Float64,1} over 16 outcomes
1 0.4470340136054422
2 0.09722448979591837
3 0.09484353741496598
4 0.02383673469387755
5 0.09995918367346938
6 0.02634013605442177
7 0.02414965986394558
8 0.0031156462585034015
9 0.0987891156462585
10 0.02346938775510204
11 0.025904761904761906
12 0.0031020408163265306
13 0.022965986394557825
14 0.0030068027210884353
15 0.002993197278911565
16 0.0032653061224489797Histograms
The histogram function counts the occurrences of each microstate identified during sampling. It is called internally by distribution , which converts the resulting Counts into Probabilities.
RecurrenceMicrostatesAnalysis.histogram — Function
histogram(core::RMACore, [x], [y])Compute the histogram of recurrence microstates for the input data [x] and [y] using the specified backend core, which must be an RMACore.
This function executes the full backend pipeline: sampling the recurrence space, constructing microstates, and evaluating recurrences.
The result is returned as a Counts object, where each index corresponds to the decimal representation of the associated microstate.
histogram(core::StandardCPUCore, [x], [y]; kwargs...)Compute the histogram of recurrence microstates for an abstract recurrence structure constructed from the input data [x] and [y].
If [x] and [y] are identical, the result corresponds to a Recurrence Plot (RP); otherwise, it corresponds to a Cross-Recurrence Plot (CRP).
The result is returned as a Counts object representing the histogram of recurrence microstates for the given input data.
This method implements the CPU backend using a CPUCore, specifically the StandardCPUCore implementation.
Arguments
core: AStandardCPUCoredefining the CPU backend configuration.[x]: Input data provided as aStateSpaceSetor anAbstractArray.[y]: Input data provided as aStateSpaceSetor anAbstractArray.
StateSpaceSet and AbstractArray inputs use different internal backends and therefore different histogram implementations. Both interfaces share the same method signature, differing only in the input data representation.
Keyword Arguments
threads: Number of threads used to compute the histogram. By default, this is set toThreads.nthreads(), which can be specified at Julia startup using--threads Nor via theJULIA_NUM_THREADSenvironment variable.
Examples
- Time series:
ssset = StateSpaceSet(rand(Float64, (1000)))
core = CPUCore(Rect(Standard(0.27), 2), SRandom(0.05))
dist = histogram(core, ssset, ssset)- Spatial data:
spatialdata = rand(Float64, (3, 50, 50))
core = CPUCore(Rect(Standard(0.5), (2, 2, 1, 1)), SRandom(0.05))
dist = histogram(core, spatialdata, spatialdata)histogram(core::StandardGPUCore, [x], [y]; kwargs...)Compute the histogram of recurrence microstates for an abstract recurrence structure constructed from the input data [x] and [y].
If [x] and [y] are identical, the result corresponds to a Recurrence Plot (RP); otherwise, it corresponds to a Cross-Recurrence Plot (CRP).
The result is returned as a Counts object representing the histogram of recurrence microstates for the given input data.
This method implements the GPU backend using a GPUCore, specifically the StandardGPUCore implementation.
Arguments
core: AStandardGPUCoredefining the GPU backend configuration.[x]: Input data provided as anAbstractGPUVector.[y]: Input data provided as anAbstractGPUVector.
Keyword Arguments
groupsize: Number of threads per GPU workgroup.
Examples
using CUDA
gpudata = StateSpaceSet(Float32.(data)) |> CuVector
core = GPUCore(CUDABackend(), Rect(Standard(0.27f0; metric = GPUEuclidean()), 2), SRandom(0.05))
dist = histogram(core, gpudata, gpudata)