Flocking model
The flock model illustrates how flocking behavior can emerge when each bird follows three simple rules:
- maintain a minimum distance from other birds to avoid collision
- fly towards the average position of neighbors
- fly in the average direction of neighbors
Defining the core structures
We begin by calling the required packages and defining an agent type representing a bird.
using Agents
using Random, LinearAlgebra
@agent struct Bird(ContinuousAgent{2, Float64})
const speed::Float64
const cohere_factor::Float64
const separation::Float64
const separate_factor::Float64
const match_factor::Float64
const visual_distance::Float64
endThe fields id and pos, which are required for agents on ContinuousSpace, are part of the struct. The field vel, which is also added by using ContinuousAgent is required for using move_agent! in ContinuousSpace with a time-stepping method. speed defines how far the bird travels in the direction defined by vel per step. separation defines the minimum distance a bird must maintain from its neighbors. visual_distance refers to the distance a bird can see and defines a radius of neighboring birds. The contribution of each rule defined above receives an importance weight: cohere_factor is the importance of maintaining the average position of neighbors, match_factor is the importance of matching the average trajectory of neighboring birds, and separate_factor is the importance of maintaining the minimum distance from neighboring birds.
The function initialize_model generates birds and returns a model object using default values.
function initialize_model(;
n_birds = 100,
speed = 1.0,
cohere_factor = 0.1,
separation = 2.0,
separate_factor = 0.25,
match_factor = 0.04,
visual_distance = 5.0,
extent = (100, 100),
seed = 42,
)
space2d = ContinuousSpace(extent; spacing = visual_distance / 1.5)
rng = Random.MersenneTwister(seed)
model = StandardABM(Bird, space2d; rng, agent_step!, container = Vector, scheduler = Schedulers.Randomly())
for _ in 1:n_birds
vel = rand(abmrng(model), SVector{2}) * 2 .- 1
add_agent!(
model,
vel,
speed,
cohere_factor,
separation,
separate_factor,
match_factor,
visual_distance,
)
end
return model
endinitialize_model (generic function with 1 method)Defining the agent_step!
agent_step! is the primary function called for each step and computes velocity according to the three rules defined above.
function agent_step!(bird, model)
# Obtain the ids of neighbors within the bird's visual distance
neighbor_agents = nearby_agents(bird, model, bird.visual_distance)
N = 0
match = separate = cohere = SVector{2}(0.0, 0.0)
# Calculate behaviour properties based on neighbors
for neighbor in neighbor_agents
N += 1
heading = get_direction(bird.pos, neighbor.pos, model)
# `cohere` computes the average position of neighboring birds
cohere += heading
# `match` computes the average trajectory of neighboring birds
match += neighbor.vel
if sum(heading .^ 2) < bird.separation^2
# `separate` repels the bird away from neighboring birds
separate -= heading
end
end
# Normalise results based on model input and neighbor count
cohere *= bird.cohere_factor
separate *= bird.separate_factor
match *= bird.match_factor
# Compute velocity based on rules defined above
bird.vel += (cohere + separate + match) / max(N, 1)
bird.vel /= norm(bird.vel)
# Move bird according to new velocity and speed
return move_agent!(bird, model, bird.speed)
end
model = initialize_model()StandardABM with 100 agents of type Bird
agents container: Vector
space: periodic continuous space with [100.0, 100.0] extent and spacing=3.3333333333333335
scheduler: Agents.Schedulers.RandomlyPlotting the flock
using CairoMakieThe great thing about abmplot is its flexibility. We can incorporate the direction of the birds when plotting them, by making the "marker" function agent_marker create a Polygon: a triangle with same orientation as the bird's velocity. It is as simple as defining the following function:
const bird_polygon = Makie.Polygon(Point2f[(-1, -1), (2, 0), (-1, 1)])
function bird_marker(b::Bird)
φ = atan(b.vel[2], b.vel[1]) #+ π/2 + π
return rotate_polygon(bird_polygon, φ)
endbird_marker (generic function with 1 method)Where we have used the utility functions scale_polygon and rotate_polygon to act on a predefined polygon. translate_polygon is also available. We now give bird_marker to abmplot, and notice how the agent_size keyword is meaningless when using polygons as markers.
model = initialize_model()
figure, = abmplot(model; agent_marker = bird_marker)
figure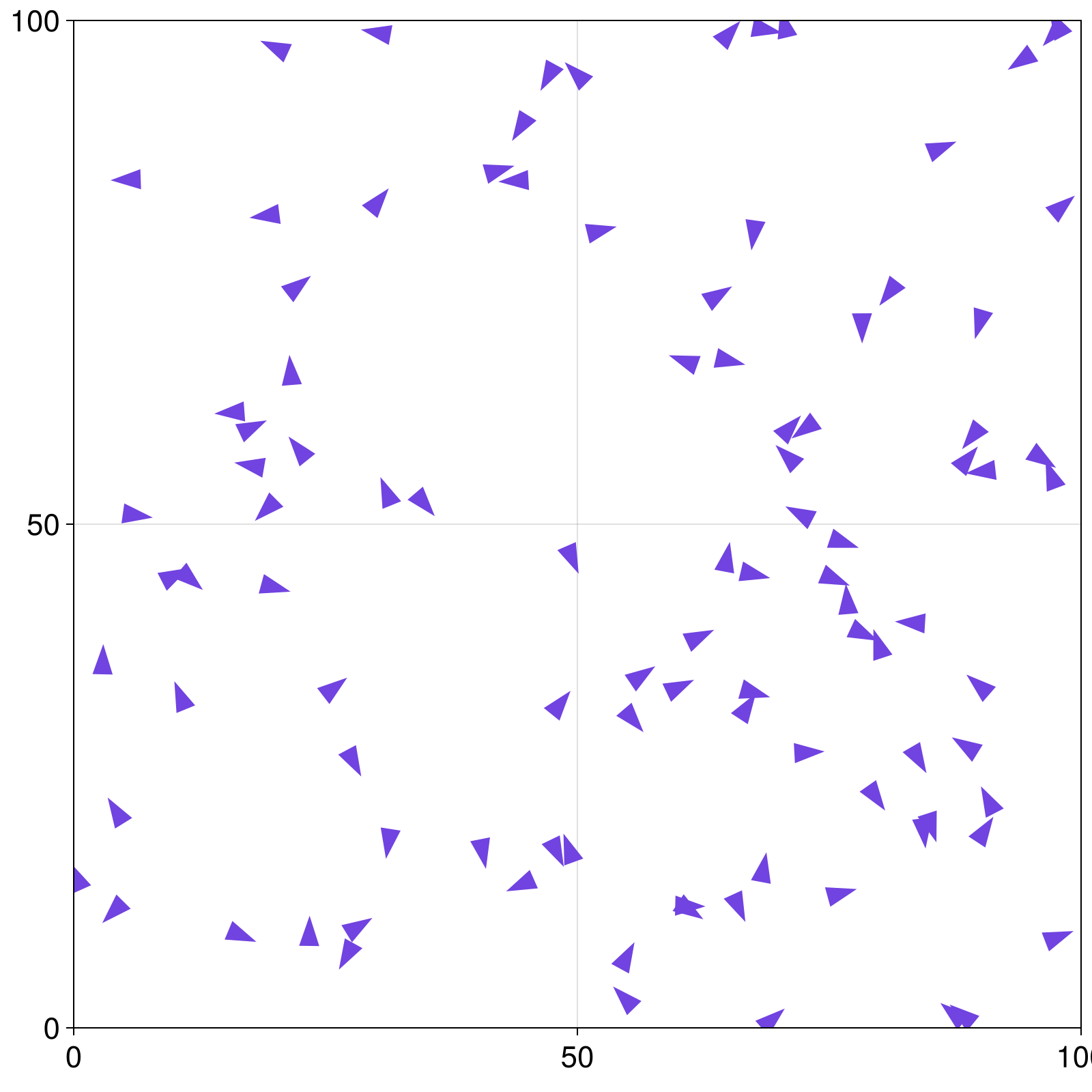
And let's also do a nice little video for it:
abmvideo(
"flocking.mp4", model;
agent_marker = bird_marker,
framerate = 20, frames = 150,
title = "Flocking"
)